Research conducted by:
Costas Maranas, the Donald B. Broughton Professor of Chemical Engineering
Tags:
chemical engineering industrial acid production modeling
Research Summary:
Microbes and other microscopic organisms could serve as sustainable “factories” to create many types of industrial materials because they naturally convert nutrients such as sugars into byproducts. However, creating industrial amounts of organic acids from renewable resources poses a challenge, because not many organisms can grow in highly acidic environments. With the help of gene editing and computational modeling tools, a team of researchers explored one type of yeast that could survive in the harsh environment created by acidic products. The team, which includes Penn State, the University of Illinois at Urbana-Champaign and Princeton University researchers, studied the yeast strain Issatchenkia orientalis, considered to be a “non-model” yeast because it has not been researched extensively. After reconstructing the yeast’s metabolism into a network model, the team examined its growth on different feedstock and subsequent byproducts. This strain was able to produce succinic acid, which is a precursor for industrial polymer production.
How Roar played a role in this research:
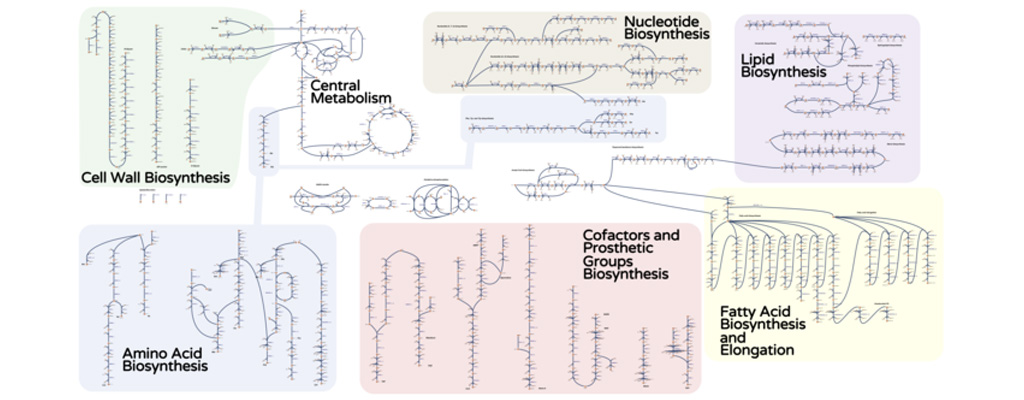
The computation-heavy approach, which ran on Penn State’s Roar supercomputer, was important in helping to refine the research direction, according to the researchers. The I. orientalis model covers 850 genes and contains 1,826 metabolic reactions, so identifying the right combination of genes and reactions to modify in order to produce succinic acid becomes a needle-in-a-haystack type of problem. Running thousands of computer simulations sifts through the hay and provides a much narrower set of experiments to test in a lab.
|
Article Title: |
Genome-scale metabolic reconstruction of the non-model yeast Issatchenkia orientalis SD108 and its application to organic acids production |
|---|---|
|
Published In: |
Metabolic Engineering Communications |
|
Abstract: |
Many platform chemicals can be produced from renewable biomass by microorganisms, with organic acids making up a large fraction. Intolerance to the resulting low pH growth conditions, however, remains a challenge for the industrial production of organic acids by microorganisms. Issatchenkia orientalis SD108 is a promising host for industrial production because it is tolerant to acidic conditions as low as pH 2.0. With the goal to systematically assess the metabolic capabilities of this non-model yeast, we developed a genome-scale metabolic model for I. orientalis SD108 spanning 850 genes, 1826 reactions, and 1702 metabolites. In order to improve the model’s quantitative predictions, organism-specific macromolecular composition and ATP maintenance requirements were determined experimentally and implemented. We examined its network topology, including essential genes and flux coupling analysis and drew comparisons with the Yeast 8.3 model for Saccharomyces cerevisiae. We explored the carbon substrate utilization and examined the organism’s production potential for the industrially-relevant succinic acid, making use of the OptKnock framework to identify gene knockouts which couple production of the targeted chemical to biomass production. The genome-scale metabolic model iIsor850 is a data-supported curated model which can inform genetic interventions for overproduction. View article on publisher's website |