Mechanical force controls the speed of protein synthesis
Posted on May 16, 2018
UNIVERSITY PARK, PA. — As cells create proteins, the proteins modulate synthesis speed by exerting a mechanical force on the molecular machine that makes them, according to a team of scientists who used a combination of computational and experimental techniques to understand this force.
Proteins power many of a cell’s vital functions, from providing structure to delivering information to fighting viruses. Defective protein synthesis is linked to numerous diseases, including subtypes of hemophilia, lung carcinoma, and cervical and vulvar cancers.
“What has been observed in the past decade is that if you change the speed at which a protein gets synthesized, you can alter how the protein behaves,” said Edward O’Brien, assistant professor of chemistry and an Institute for Computational and Data Sciences co-hire, Penn State. “We tried to identify new factors that influence protein synthesis speed.”
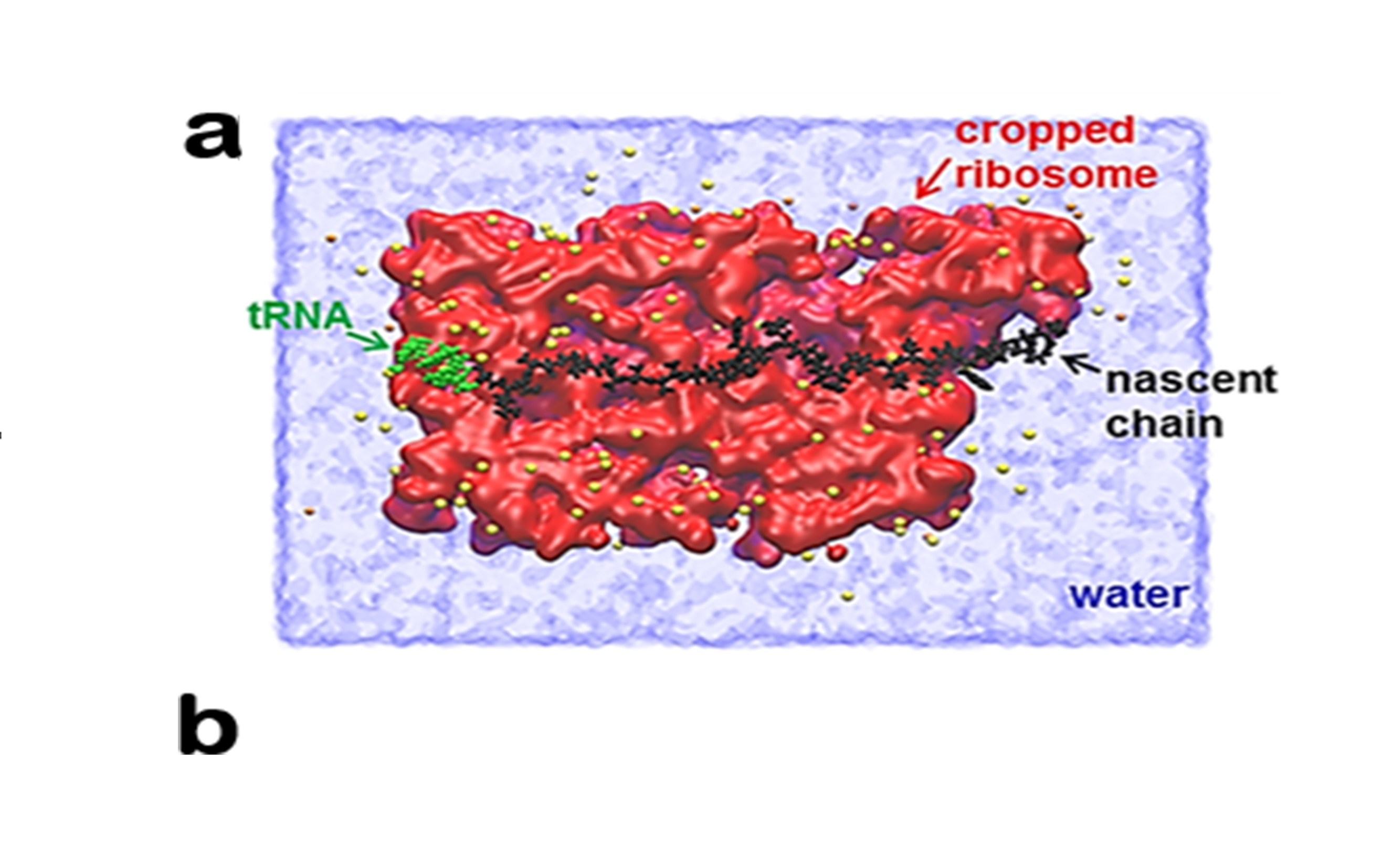
Ribosomes, tiny factories in the cell, stitch amino acids into a long chain to create proteins. During this process, newly synthesized protein segments pass through a narrow tunnel of the ribosome. When they exit the tunnel, proteins naturally pull away from the ribosome, O’Brien said.
“These protein molecules like to be in regions of space with a lot of free volume where they can move around, rather than in confined narrow spaces,” said O’Brien.
The force that pulls the protein from the ribosome is an entropic pulling force that happens naturally, according to O’Brien. Entropic force in a system is a force resulting from the entire system’s tendency to increase its entropy, rather than from a particular underlying microscopic force. Entropy is the tendency for systems to become more disordered over time.
“That pulling force gets propagated back to where the synthesis is occurring within the ribosome, and modulates that process,” said O’Brien.
The researchers observed that unstructured protein segments generate piconewtons of force and that this force is transmitted across the ribosome molecular machine, and that it affects the speed at which amino acids are stitched together.
The team started their study from experimental measurements that detected how much proteins stretched on the ribosome. The researchers input this information into high-performance computer simulations that ran for months using both the Penn State Institute for Computational and Data Sciences’ Advanced CyberInfrastructure and the Extreme Science and Engineering Discovery Environment, an NSF-funded virtual organization. These simulations allowed them to see how protein synthesis was carried out under numerous conditions.
“By understanding the factors governing the speed of protein synthesis, we can now start to understand how protein synthesis affects downstream processes involving protein structure and function, including various diseases,” said O’Brien.
The researchers published their findings in the Journal of the American Chemical Society.
Collaborators on this work include recent graduate Benjamin Fritch, and graduate student in chemistry Daniel Nissley, Penn State; Andrey Kosolapov and Carol Deutsch, University of Pennsylvania; and Phillip Hudson and H. Lee Woodcock, University of South Florida.
The National Science Foundation, Penn State and the University of South Florida supported this work.
Image: Benjamin Fritch
Share
Related Posts
- Professor receives NSF grant to model cell disorder in heart
- Featured Researcher: Nick Tusay
- Multi-institutional team to use AI to evaluate social, behavioral science claims
- NSF invests in cyberinfrastructure institute to harness cosmic data
- Center for Immersive Experiences set to debut, serving researchers and students
- Distant Suns, Distant Worlds
- CyberScience Seminar: Researcher to discuss how AI can help people avoid adverse drug interactions
- AI could offer warnings about serious side effects of drug-drug interactions
- Taking RTKI drugs during radiotherapy may not aid survival, worsens side effects
- Cost-effective cloud research computing options now available for researchers
- Costs of natural disasters are increasing at the high end
- Model helps choose wind farm locations, predicts output
- Virus may jump species through ‘rock-and-roll’ motion with receptors
- Researchers seek to revolutionize catalyst design with machine learning
- Resilient Resumes team places third in Nittany AI Challenge
- ‘AI in Action’: Machine learning may help scientists explore deep sleep
- Clickbait Secrets Exposed! Humans and AI team up to improve clickbait detection
- Focusing computational power for more accurate, efficient weather forecasts
- How many Earth-like planets are around sun-like stars?
- SMH! Brains trained on e-devices may struggle to understand scientific info
- Whole genome sequencing may help officials get a handle on disease outbreaks
- New tool could reduce security analysts’ workloads by automating data triage
- Careful analysis of volcano’s plumbing system may give tips on pending eruptions
- Reducing farm greenhouse gas emissions may plant the seed for a cooler planet
- Using artificial intelligence to detect discrimination
- Four ways scholars say we can cut the chances of nasty satellite data surprises
- Game theory shows why stigmatization may not make sense in modern society
- Older adults can serve communities as engines of everyday innovation
- Pig-Pen effect: Mixing skin oil and ozone can produce a personal pollution cloud
- Researchers find genes that could help create more resilient chickens
- Despite dire predictions, levels of social support remain steady in the U.S.
- For many, friends and family, not doctors, serve as a gateway to opioid misuse
- New algorithm may help people store more pictures, share videos faster
- Head named for Ken and Mary Alice Lindquist Department of Nuclear Engineering
- Scientific evidence boosts action for activists, decreases action for scientists
- People explore options, then selectively represent good options to make difficult decisions
- Map reveals that lynching extended far beyond the deep South
- Gravitational forces in protoplanetary disks push super-Earths close to stars
- Supercomputer cluster donation helps turn high school class into climate science research lab
- Believing machines can out-do people may fuel acceptance of self-driving cars
- People more likely to trust machines than humans with their private info
- IBM donates system to Penn State to advance AI research
- ICS Seed Grants to power projects that use AI, machine learning for common good
- Penn State Berks team advances to MVP Phase of Nittany AI Challenge
- Creepy computers or people partners? Working to make AI that enhances humanity
- Sky is clearing for using AI to probe weather variability
- ‘AI will see you now’: Panel to discuss the AI revolution in health and medicine
- Privacy law scholars must address potential for nasty satellite data surprises
- Researchers take aim at hackers trying to attack high-value AI models
- Girls, economically disadvantaged less likely to get parental urging to study computers
- Seed grants awarded to projects using Twitter data
- Researchers find features that shape mechanical force during protein synthesis
- A peek at living room decor suggests how decorations vary around the world
- Interactive websites may cause antismoking messages to backfire
- Changing how government assesses risk may ease fallout from extreme financial events
- Penn State’s Leadership in AI Research
- ICS co-sponsors Health, Environment Seed Grant Program
- Symposium at U.S. Capitol seeks solutions to election security
